I’m pretty sure most readers of this blog are already familiar with Rmarkdown and Github. In this post I don’t pretend to invent the wheel but rather give a quick run-down of how I set-up and use these tools to produce high quality and scalable (in human time) reproducible data science development code.
Github
While data science processes usually don’t involve the exact same workflows like software development (for which Git was originally intended) I think Git is actually very well suited to the iterative nature of data-science tasks.
When walking down different avenues in the exploration path, it’s worth while to have them reside in different branches. That way instead of jotting down in general pointers what you did along with some code snippets in some text file (or god-forbid word when you want to have images as well) you can instead go back to the relevant branch, see the different iterations and read a neat report with code and images. You can even re-visit ideas that didn’t make it into the master branch. Be sure to use informative branch names and commit messages!
Below is in illustration of how that process might look like:
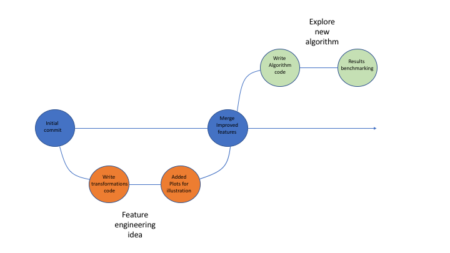
Using Github allows one to easily package his code, supporting files etc (using repos) and share it with fellow researches, which can in turn clone the repo, re-run the code and go through all the development iterations without a hassle.
Rmarkdown
Most people familiar with Rmarkdown know it’s a great tool to write neat reports in all sorts of formats (html, PDF and even word!). One format that really makes it a great combo with Github is the github_document format. While one can’t view HTML files on Github, the output file from a github_document knit is an .md file which renders perfectly well on github, supporting images, tables, math, table of contents and many other.
What some may not realize is that Rmarkdown is also a great development tool in itself. It behaves much like the popular Jupiter notebooks, with plots, tables and equations showing next to the code that generated them. What’s more, it has tons of cool features that really support reproducible development such as:
- The first r-chunk (labled “setup” in the Rstudio template) always runs once when you execute code within chunks following it (pressing ctrl+Enter). It’s handy to load all packages used in later chucks (I like installing missing ones too) in this chunk such that whenever you run code within any of the chunks below it the needed packages are loaded.
- When running code from within a chunk (pressing ctrl+Enter) the working directory will always be the one which the .Rmd file is located at. In short this means no more worrying about setting the working directory – be it when working on several projects simultaneously or when cloning a repo from Github.
- It has many cool code execution tools such as a button to run code in all chunks up to the current one, run all code in the current chunk and it has a green progress bar so you don’t get lost too!
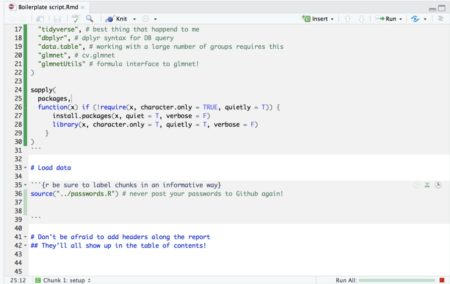
- If your script is so long that scrolling around it becomes tedious, you can use this neat feature in Rstudio: When viewing Rmarkdown files you can view an interactive table of contents that enables you to jump between sections (defined by # headers) in your code:
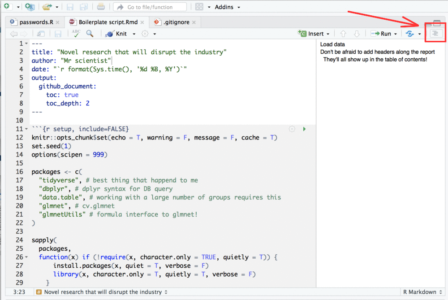
To summarize this section, I would highly recommend developing with Rmd files rather than R files.
A few set-up tips
- Place a file “passwords.R” with all passwords in the directory to which you clone repos and source it via the Rmd. That way you don’t accidentally publish your passwords to Github
- I like working with cache on all chunks in my Rmd. It’s usually good practice to avoid uploading the cache files generated in the process to Github so be sure to add to your .gitignore file the file types: *.RData, *.rdb, *.rdx, *.rds, *__packages
- Github renders CSV files pretty nicely (and enables searching them conveniently) so if you have some reference tables you want to include and you have a *.csv entry in your .gitignore file, you may want to add to your .gitignore the following entry: !reference_table_which_renders_nicely_on_github.csv to exclude it from the exclusion list.
Sample Reproducible development repo
Feel free to clone the sample reproducible development repo below and get your reproducible project running ASAP!
https://github.com/IyarLin/boilerplate-script