As the post on “hello world” functions has been quite appreciated by the R community, here follows the second round of functions for wannabe R programmer.
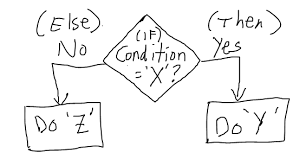
# If else statement:
# See the code syntax below for if else statement
x=10
if(x>1){
print(“x is greater than 1”)
}else{
print(“x is less than 1”)
}
# See the code below for nested if else statement
x=10
if(x>1 & x<7){
print(“x is between 1 and 7”)} else if(x>8 & x< 15){
print(“x is between 8 and 15”)
}
# For loops:
# Below code shows for loop implementation
x = c(1,2,3,4,5)
for(i in 1:5){
print(x[i])
}
# While loop :
# Below code shows while loop in R
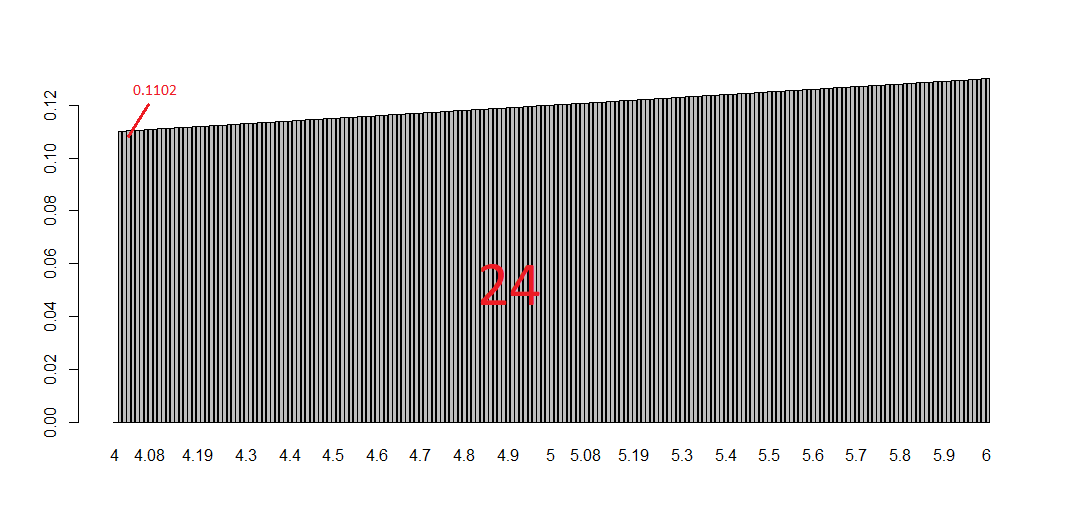
x = 2.987
while(x <= 4.987) {
x = x + 0.987
print(c(x,x-2,x-1))
}
# Repeat Loop:
# The repeat loop is an infinite loop and used in association with a break statement.
# Below code shows repeat loop:
a = 1
repeat{
print(a)
a = a+1
if (a > 4) {
break
}
}
# Break statement:
# A break statement is used in a loop to stop the iterations and flow the control outside of the loop.
#Below code shows break statement:
x = 1:10
for (i in x){
if (i == 6){
break
}
print(i)
}
# Next statement:
# Next statement enables to skip the current iteration of a loop without terminating it.
#Below code shows next statement
x = 1: 4
for (i in x) {
if (i == 2){
next
}
print(i)
}
# function
words = c(“R”, “datascience”, “machinelearning”,”algorithms”,”AI”)
words.names = function(x) {
for(name in x){
print(name)
}
}
words.names(words) # Calling the function
# extract the elements above the main diagonal of a (square) matrix
# example of a correlation matrix
cor_matrix <- matrix(c(1, -0.25, 0.89, -0.25, 1, -0.54, 0.89, -0.54, 1), 3,3)
rownames(cor_matrix) <- c(“A”,”B”,”C”)
colnames(cor_matrix) <- c(“A”,”B”,”C”)
cor_matrix
rho <- list()
name <- colnames(cor_matrix)
var1 <- list()
var2 <- list()
for (i in 1:ncol(cor_matrix)){
for (j in 1:ncol(cor_matrix)){
if (i != j & i<j){
rho <- c(rho,cor_matrix[i,j])
var1 <- c(var1, name[i])
var2 <- c(var2, name[j])
}
}
}
d <- data.frame(var1=as.character(var1), var2=as.character(var2), rho=as.numeric(rho))
d
var1 var2 rho
1 A B -0.25
2 A C 0.89
3 B C -0.54
As programming is the best way to learn and think, have fun programming awesome functions! This post is also shared in R-bloggers and LinkedIn


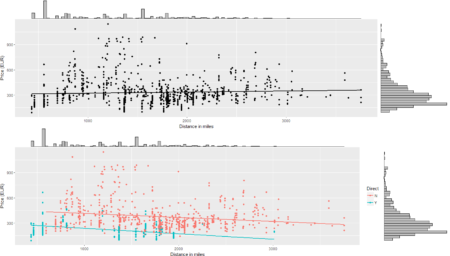 To conclude the 4-step (flight) trip from data acquisition to data analysis, let's recap the most important concepts described in each of the post:
1)
To conclude the 4-step (flight) trip from data acquisition to data analysis, let's recap the most important concepts described in each of the post:
1) 
 The collection of example flight data in json format available in
The collection of example flight data in json format available in 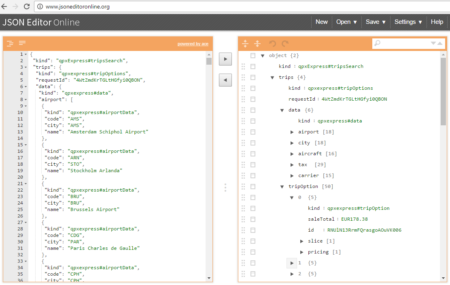 However, looking deeply into the object, several other elements are provided as the distance in mile, the segment, the duration, the carrier, etc. The R parser to transform the json structure in a usable dataframe requires the dplyr library for using the pipe operator (%>%) to streamline the code and make the parser more readable. Nevertheless, the library actually wrangling through the lines is tidyjson and its powerful functions:
However, looking deeply into the object, several other elements are provided as the distance in mile, the segment, the duration, the carrier, etc. The R parser to transform the json structure in a usable dataframe requires the dplyr library for using the pipe operator (%>%) to streamline the code and make the parser more readable. Nevertheless, the library actually wrangling through the lines is tidyjson and its powerful functions: