In this post, taken from the book R Data Mining by Andrea Cirillo, we’ll be looking at how to scrape PDF files using R. It’s a relatively straightforward way to look at text mining – but it can be challenging if you don’t know exactly what you’re doing.
In this post, taken from the book R Data Mining by Andrea Cirillo, we’ll be looking at how to scrape PDF files using R. It’s a relatively straightforward way to look at text mining – but it can be challenging if you don’t know exactly what you’re doing.Until January 15th, every single eBook and video by Packt is just $5! Start exploring some of Packt’s huge range of R titles here.
You may not be aware of this, but some organizations create something called a ‘customer card’ for every single customer they deal with. This is quite an informal document that contains some relevant information related to the customer, such as the industry and the date of foundation. Probably the most precious information contained within these cards is the comments they write down about the customers. Let me show you one of them:
 My plan was the following—get the information from these cards and analyze it to discover whether some kind of common traits emerge.
My plan was the following—get the information from these cards and analyze it to discover whether some kind of common traits emerge.As you may already know, at the moment this information is presented in an unstructured way; that is, we are dealing with unstructured data. Before trying to analyze this data, we will have to gather it in our analysis environment and give it some kind of structure.
Technically, what we are going to do here is called text mining, which generally refers to the activity of gaining knowledge from texts. The techniques we are going to employ are the following:
- Sentiment analysis
- Wordclouds
- N-gram analysis
- Network analysis
Getting a list of documents in a folder
First of all, we need to get a list of customer cards we were from the commercial department. I have stored all of them within the ‘data’ folder on my workspace. Let’s use list.files() to get them:file_vector <- list.files(path = "data")Nice! We can inspect this looking at the head of it. Using the following command:
file_vector %>% head() [1] "banking.xls" "Betasoloin.pdf" "Burl Whirl.pdf" "BUSINESSCENTER.pdf" [5] "Buzzmaker.pdf" "BuzzSaw Publicity.pdf"Uhm… not exactly what we need. I can see there are also .xls files. We can remove them using the grepl() function, which performs partial matches on strings, returning TRUE if the pattern required is found, or FALSE if not. We are going to set the following test here: give me TRUE if you find .pdf in the filename, and FALSE if not:
grepl(".pdf",file_list)
[1] FALSE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE FALSE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[18] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[35] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[52] TRUE TRUE TRUE TRUE FALSE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[69] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[86] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[103] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE FALSE FALSE TRUE
[120] TRUE
As you can see, the first match results in a FALSE since it is related to the .xls file we saw before. We can now filter our list of files by simply passing these matching results to the list itself. More precisely, we will slice our list, selecting only those records where our grepl() call returns TRUE:
pdf_list <- file_vector[grepl(".pdf",file_list)]
Did you understand [grepl(“.pdf”,file_list)] ? It is actually a way to access one or more indexes within a vector, which in our case are the indexes corresponding to “.pdf”, exactly the same as we printed out before.
If you now look at the list, you will see that only PDF filenames are shown on it.
Reading PDF files into R via pdf_text()
R comes with a really useful that’s employed tasks related to PDFs. This is named pdftools, and beside the pdf_text function we are going to employ here, it also contains other relevant functions that are used to get different kinds of information related to the PDF file into R. For our purposes, it will be enough to get all of the textual information contained within each of the PDF files. First of all, let’s try this on a single document; we will try to scale it later on the whole set of documents. The only required argument to make pdf_text work is the path to the document. The object resulting from this application will be a character vector of length 1:pdf_text("data/BUSINESSCENTER.pdf")
[1] "BUSINESSCENTER business profile\nInformation below are provided under non disclosure agreement. date of enquery: 12.05.2017\ndate of foundation: 1993-05-18\nindustry: Non-profit\nshare holders: Helene Wurm ; Meryl Savant ; Sydney Wadley\ncomments\nThis is one of our worst customer. It really often miss payments even if for just a couple of days. We have\nproblems finding useful contact persons. The only person we can have had occasion to deal with was the\nfiscal expert, since all other relevant person denied any kind of contact.\n 1\n"
If you compare this with the original PDF document you can easily see that all of the information is available even if it is definitely not ready to be analyzed. What do you think is the next step needed to make our data more useful?
We first need to split our string into lines in order to give our data a structure that is closer to the original one, that is, made of paragraphs. To split our string into separate records, we can use the strsplit() function, which is a base R function. It requires the string to be split and the token that decides where the string has to be split as arguments. If you now look at the string, you’ll notice that where we found the end of a line in the original document, for instance after the words business profile, we now find the \n token. This is commonly employed in text formats to mark the end of a line.
We will therefore use this token as a split argument:
pdf_text("data/BUSINESSCENTER.pdf") %>% strsplit(split = "\n")
[[1]]
[1] "BUSINESSCENTER business profile"
[2] "Information below are provided under non disclosure agreement. date of enquery: 12.05.2017"
[3] "date of foundation: 1993-05-18"
[4] "industry: Non-profit"
[5] "share holders: Helene Wurm ; Meryl Savant ; Sydney Wadley"
[6] "comments"
[7] "This is one of our worst customer. It really often miss payments even if for just a couple of days. We have"
[8] "problems finding useful contact persons. The only person we can have had occasion to deal with was the"
[9] "fiscal expert, since all other relevant person denied any kind of contact."
[10] " 1"
strsplit() returns a list with an element for each element of the character vector passed as argument; within each list element, there is a vector with the split string.
Isn’t that better? I definitely think it is. The last thing we need to do before actually doing text mining on our data is to apply those treatments to all of the PDF files and gather the results into a conveniently arranged data frame.
Iteratively extracting text from a set of documents with a for loop
What we want to do here is run trough the list of files and for filename found there, we run the pdf_text() function and then the strsplit() function to get an object similar to the one we have seen with our test. A convenient way to do this is by employing a ‘for’ loop. These structures basically do this to their content: Repeat this instruction n times and then stop. Let me show you a typical ‘for’ loop:for (i in 1:3){
print(i)
}
If we run this, we obtain the following:
[1] 1 [1] 2 [1] 3This means that the loop runs three times and therefore repeats the instructions included within the brackets three times. What is the only thing that seems to change every time? It is the value of i. This variable, which is usually called counter, is basically what the loop employs to understand when to stop iterating. When the loop execution starts, the loop starts increasing the value of the counter, going from 1 to 3 in our example. The for loop repeats the instructions between the brackets for each element of the values of the vector following the in clause in the for command. At each step, the value of the variable before in (i in this case) takes one value of the sequence from the vector itself. The counter is also useful within the loop itself, and it is usually employed to iterate within an object in which some kind of manipulation is desired. Take, for instance, a vector defined like this:
vector <- c(1,2,3)Imagine we want to increase the value of every element of the vector by 1. We can do this by employing a loop such as this:
for (i in 1:3){
vector[i] <- vector[i]+1
}
If you look closely at the loop, you’ll realize that the instruction needs to access the element of the vector with an index equal to i and modify this value by 1. The counter here is useful because it will allow iteration on all vectors from 1 to 3.
Be aware that this is actually not a best practice because loops tend to be quite computationally expensive, and they should be employed when no other valid alternative is available. For instance, we can obtain the same result here by working directly on the whole vector, as follows:
vector_increased <- vector +1If you are interested in the topic of avoiding loops where they are not necessary, I can share with you some relevant material on this. For our purposes, we are going to apply loops to go through the pdf_list object, and apply the pdf_text function and subsequently the strsplit() function to each element of this list:
corpus_raw <- data.frame("company" = c(),"text" = c())
for (i in 1:length(pdf_list)){
print(i)
pdf_text(paste("data/", pdf_list[i],sep = "")) %>%
strsplit("\n")-> document_text
data.frame("company" = gsub(x =pdf_list[i],pattern = ".pdf", replacement = ""),
"text" = document_text, stringsAsFactors = FALSE) -> document
colnames(document) <- c("company", "text")
corpus_raw <- rbind(corpus_raw,document)
}
Let’s get closer to the loop: we first have a call to the pdf_text function, passing an element of pdf_list as an argument; it is defined as referencing the i position in the list. Once we have done this, we can move on to apply the strsplit() function to the resulting string. We define the document object, which contains two columns, in this way:
- company, which stores the name of the PDF without the .pdf token; this is the name of the company
- text, which stores the text resulting from the extraction
corpus_raw %>% head()
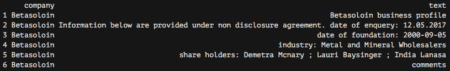
This is a well-structured object, ready for some text mining. Nevertheless, if we look closely at our PDF customer cards, we can see that there are three different kinds of information and they should be handled differently:
- Repeated information, such as the confidentiality disclosure on the second line and the date of enquiry (12.05.2017)
- Structured attributes, for instance, date of foundation or industry
- Strictly unstructured data, which is in the comments paragraph
- 12.05.2017: This denotes the line showing the non-disclosure agreement and the date of inquiry.
- business profile: This denotes the title of the document, containing the name of the company. We already have this information stored within the company column.
- comments: This is the name of the last paragraph.
- 1: This represents the number of the page and is always the same on every card.
corpus_raw %>%
filter(!grepl("12.05.2017",text)) %>%
filter(!grepl("business profile",text)) %>%
filter(!grepl("comments",text)) %>%
filter(!grepl("1",text)) -> corpus
Now that we have removed those useless things, we can actually split the data frame into two sub-data frames based on what is returned by the grepl function when searching for the following tokens, which point to the structured attributes we discussed previously:
- date of foundation
- industry
- shareholders
We are going to create two different data frames here; one is called ‘information’ and the other is called ‘comments’:
corpus %>%
filter(!grepl(c("date of foundation"),text)) %>%
filter(!grepl(c( "industry"),text)) %>%
filter(!grepl(c( "share holders"),text)) -> comments
corpus %>%
filter(grepl(("date of foundation"),text)|grepl(( "industry"),text)|grepl(( "share holders"),text))-> information
As you can see, the two data treatments are nearly the opposite of each other, since the first looks for lines showing none of the three tokens while the other looks for records showing at least one of the tokens.
Let’s inspect the results by employing the head function:
information %>% head() comments %>% head()
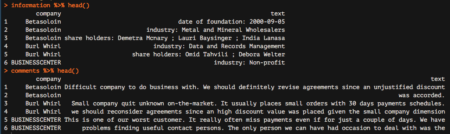 Great! We are nearly done. We are now going to start analyzing the comments data frame, which reports all comments from our colleagues. The very last step needed to make this data frame ready for subsequent analyzes is to tokenize it, which basically means separating it into different rows for all the words available within the text column. To do this, we are going to leverage the unnest_tokens function, which basically splits each line into words and produces a new row for each word, having taken care to repeat the corresponding value within the other columns of the original data frame.
This function comes from the recent tidytext package by Julia Silge and Davide Robinson, which provides an organic framework of utilities for text mining tasks. It follows the tidyverse framework, which you should already know about if you are using the dplyr package.
Let’s see how to apply the unnest_tokens function to our data:
Great! We are nearly done. We are now going to start analyzing the comments data frame, which reports all comments from our colleagues. The very last step needed to make this data frame ready for subsequent analyzes is to tokenize it, which basically means separating it into different rows for all the words available within the text column. To do this, we are going to leverage the unnest_tokens function, which basically splits each line into words and produces a new row for each word, having taken care to repeat the corresponding value within the other columns of the original data frame.
This function comes from the recent tidytext package by Julia Silge and Davide Robinson, which provides an organic framework of utilities for text mining tasks. It follows the tidyverse framework, which you should already know about if you are using the dplyr package.
Let’s see how to apply the unnest_tokens function to our data:
comments %>% unnest_tokens(word,text)-> comments_tidyIf we now look at the resulting data frame, we can see the following:
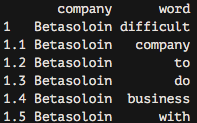 As you can see, we now have each word separated into a single record.
As you can see, we now have each word separated into a single record.Thanks for reading! We hope you enjoyed this extract taken from R Data Mining.
Explore $5 R eBooks and videos from Packt until January 15th 2018.
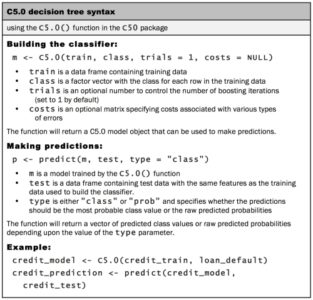

 For the first iteration of our credit approval model, we’ll use the default C5.0 configuration, as shown in the following code. The 17th column in credit_train is the default class variable, so we need to exclude it from the training data frame, but supply it as the target factor vector for classification:
For the first iteration of our credit approval model, we’ll use the default C5.0 configuration, as shown in the following code. The 17th column in credit_train is the default class variable, so we need to exclude it from the training data frame, but supply it as the target factor vector for classification:
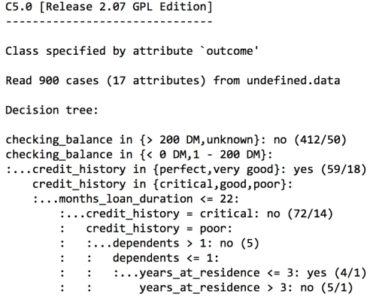 The preceding output shows some of the first branches in the decision tree. The first three lines could be represented in plain language as:
The preceding output shows some of the first branches in the decision tree. The first three lines could be represented in plain language as: